A team led by investigators at Massachusetts General Hospital (MGH) in Boston, USA, recently used deep learning to uncover insights into the genetic basis for variation in the size of the aorta. In addition to identifying at-risk individuals, the researchers believe that the findings may point to new preventive and therapeutic targets.
A team led by investigators at Massachusetts General Hospital (MGH) in Boston, USA, recently used deep learning to uncover insights into the genetic basis for variation in the size of the aorta. In addition to identifying at-risk individuals, the researchers believe that the findings may point to new preventive and therapeutic targets.
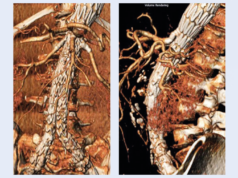
The research, which is published in Nature Genetics, relied on data from the UK Biobank, a study that performed multiple magnetic resonance imaging tests of the heart and aorta in more than 40,000 people. “There were no aortic measurements provided by the UK Biobank, and we wanted to read the aortic diameter in all of the images collected,” explains lead author James Pirruccello, a cardiologist at MGH and an instructor in medicine at Harvard Medical School (Boston, USA). “That is very hard for a human to do because it would take a long time, which motivated our use of deep learning models to do this process at a large scale.”
The researchers trained deep learning models to evaluate the dimensions of the ascending and descending sections of the aorta in 4.6 million cardiac images. They then analysed the study participants’ genes to identify variations in 82 genetic loci linked to the diameter of the ascending aorta and 47 linked to the diameter of the descending aorta. Some of the loci were near genes with known associations with aortic disease.
“When we added up the genetic variants into what is called a polygenic score, people with a higher score were more likely to be diagnosed with aortic aneurysm by a doctor,” says Pirruccello. “This suggests that, after further development and testing, such a score might one day be useful to help us identify people at high risk of an aneurysm. The genetic loci that we discovered also offer a useful starting point for trying to identify new drug targets for aortic enlargement.”
Pirruccello adds that the findings also provide supportive evidence that deep learning and other machine learning methods can help accelerate scientific analyses of complex biomedical data such as imaging results.
This work was supported by Leducq, the National Institutes of Health, the American Heart Association, the John S LaDue Memorial Fellowship, a Sarnoff Cardiovascular Research Foundation Scholar Award, the Burroughs Wellcome Fund, the Fredman Fellowship for Aortic Disease, the Toomey Fund for Aortic Dissection Research, Bayer AG, and the Susan Eid Tumor Heterogeneity Initiative.